In this tutorial, I would like to share how one can import SSR (simple sequence repeats) marker data on to TASSEL software, and perfrom analyses such as imputation, General Linear Model and Principle Component Analysis.
1.0 Installing TASSEL software
Download and install the latest version of the TASSEL software at this link:
https://www.maizegenetics.net/tassel
Ignore this step if you already have installed TASSEL.
1.1 Input data
The input SSR marker data must formated as instructed below:
- Add "Numeric” in the first line of the data
- The missing data (-1) was denoted as “NA”
- TASSEL only allows numeric marker data ranging from 0 to 1
- Save your numeric data in a text file format (.txt) prior to loading in tassel
Note: If you have two alleles per SSR marker then you can load it on TASSEL as a numeric data as 0 for major and 1 for minor alleles. However, If SSRs alleles have more than 2 alleles per marker I suggest converting as below:
- Most common = A
- 2nd most common = C
- 3rd most common = G
- 4th most common = T
- 5th most common = +
- All other alleles = - (dash)
See an example of the input file:
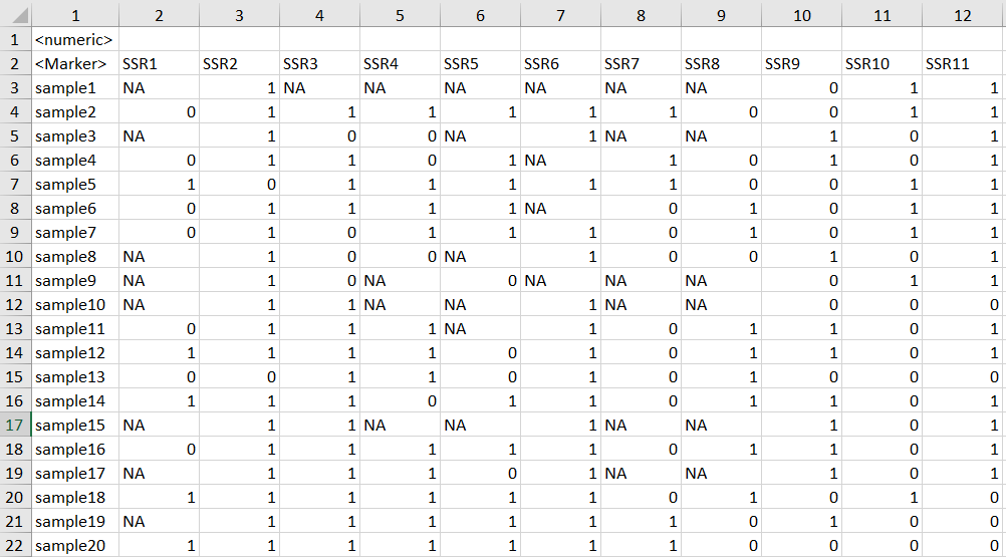
1.2 Importing phenotype and genotype files
Import the files by following the steps shown below.
Tip! Both files can be opened at same time holding CTRL and clicking the file names.
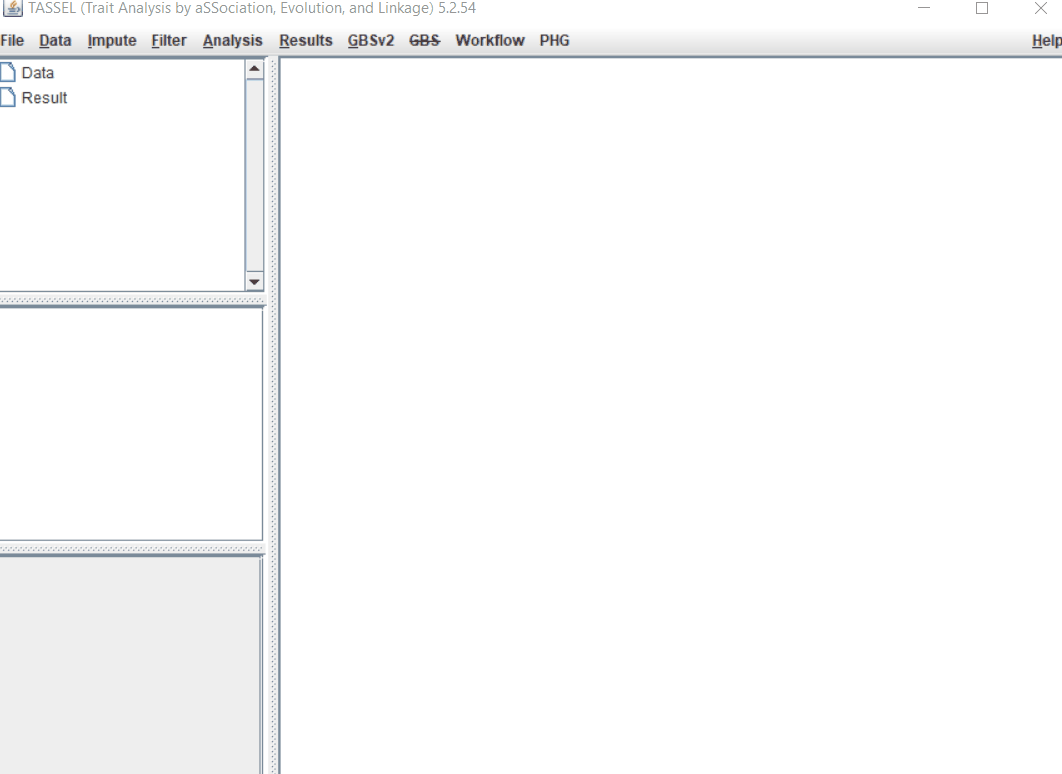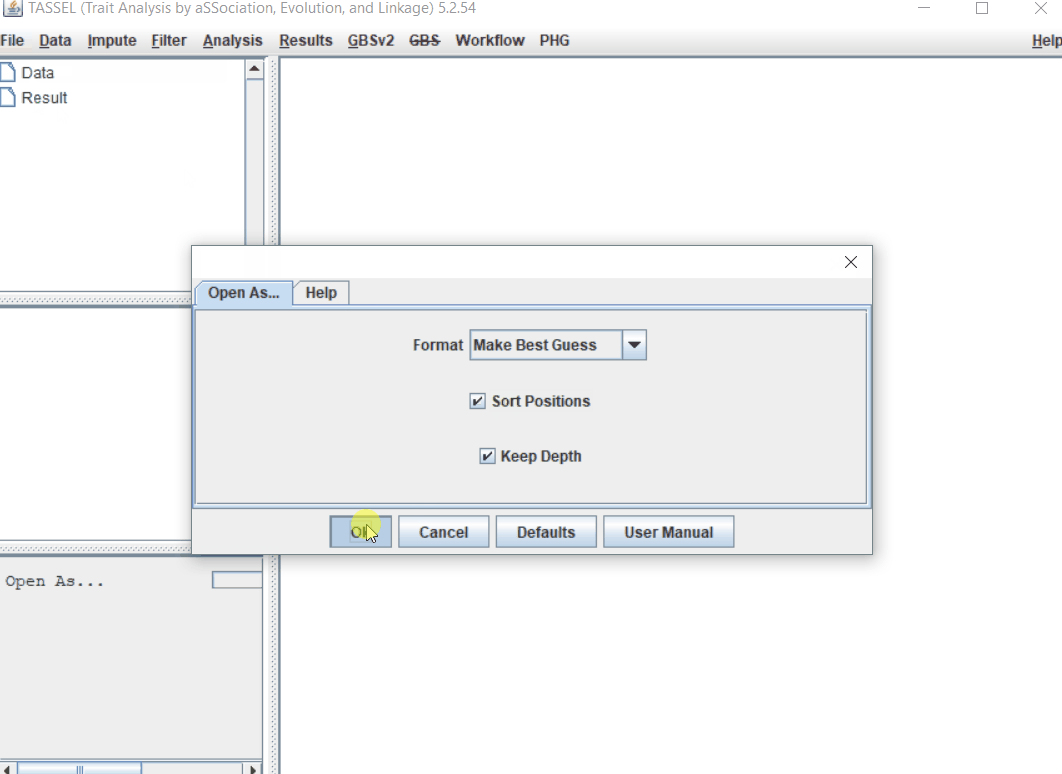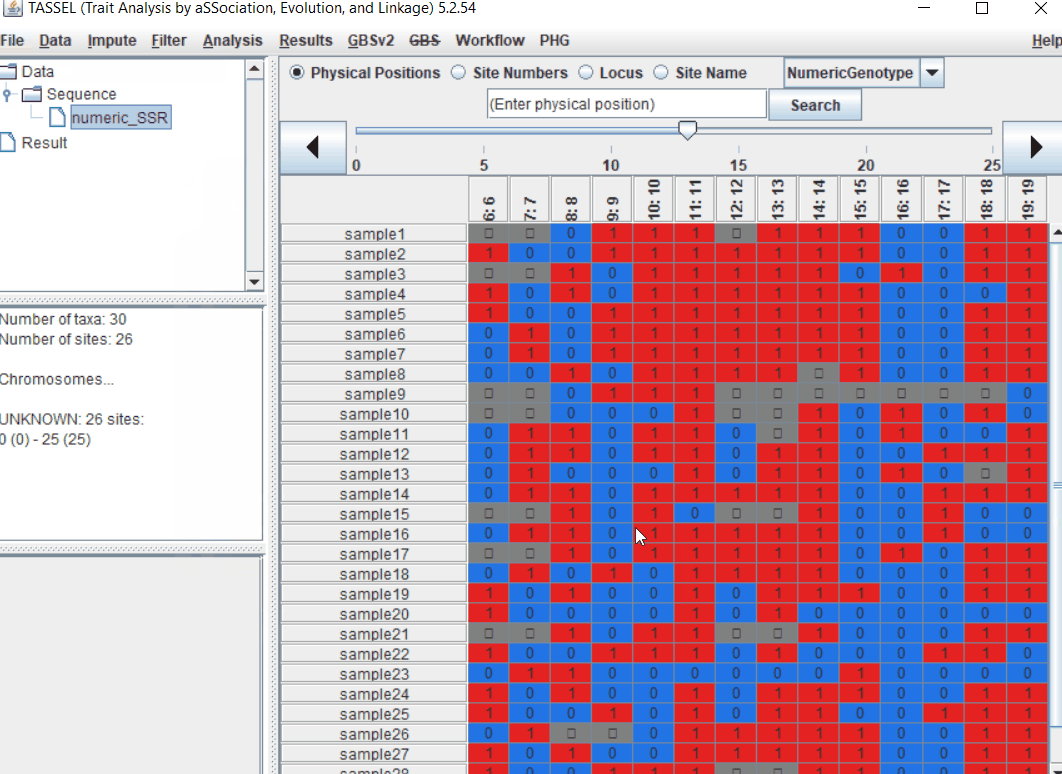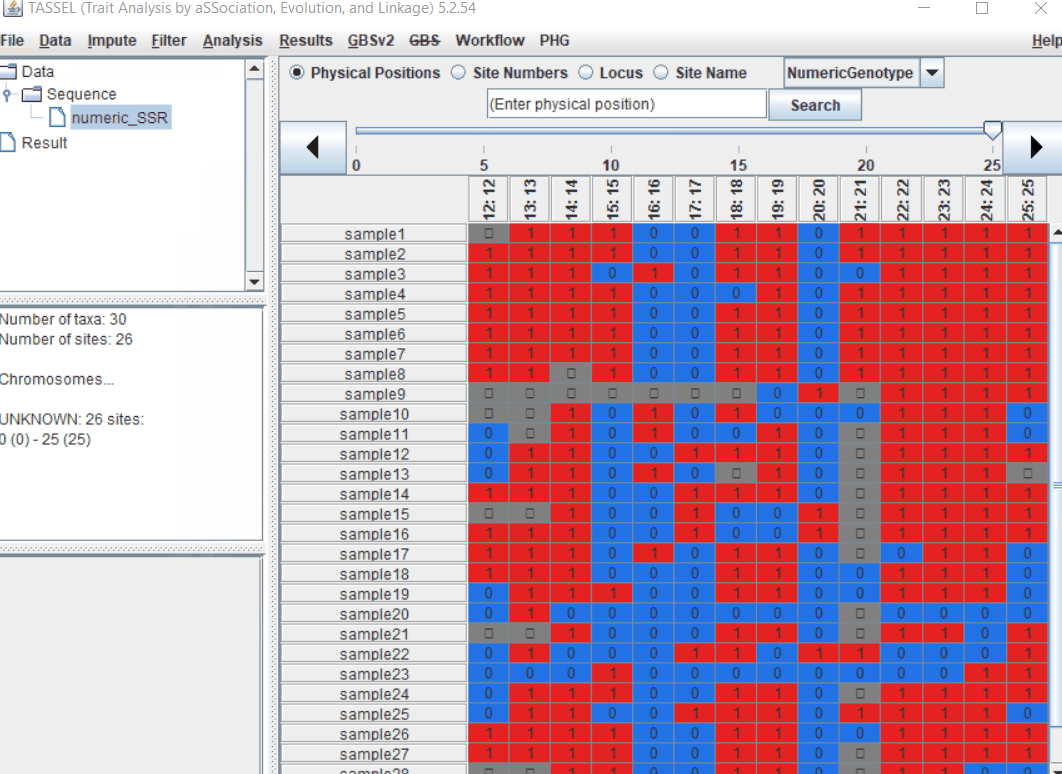
1.3 Imputation
This is an optional step, however, in order to conduct PCA analysis you need to impute and should not have any missing data.
Follow the below steps to perform numerical imputation:
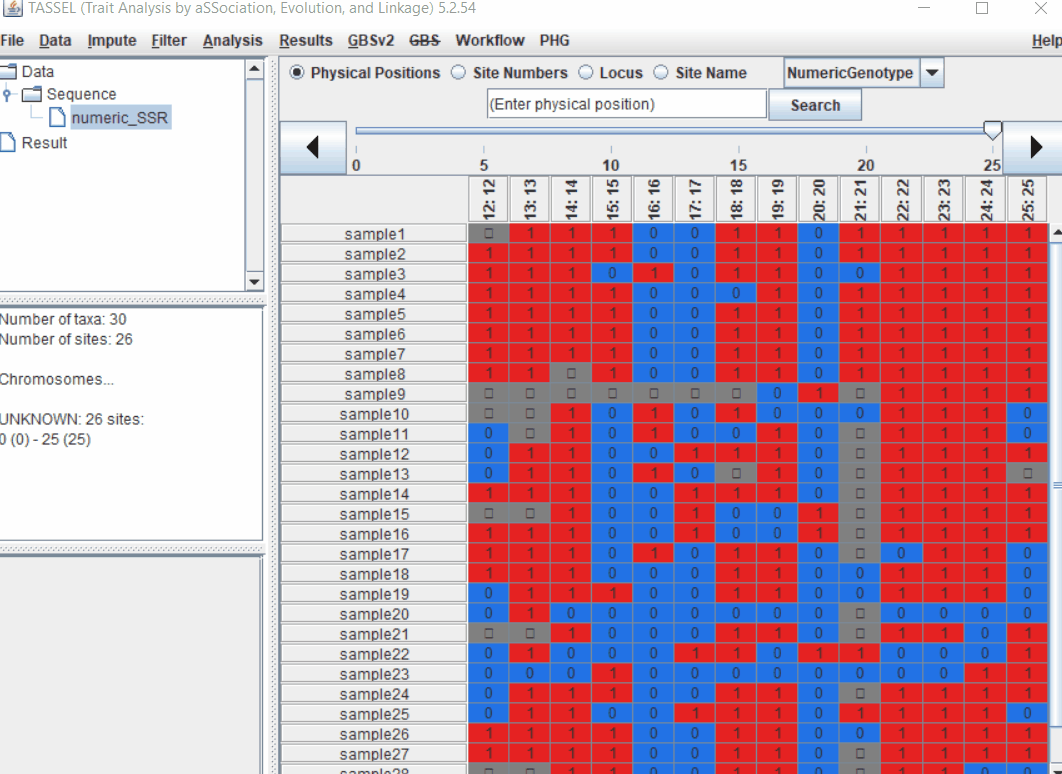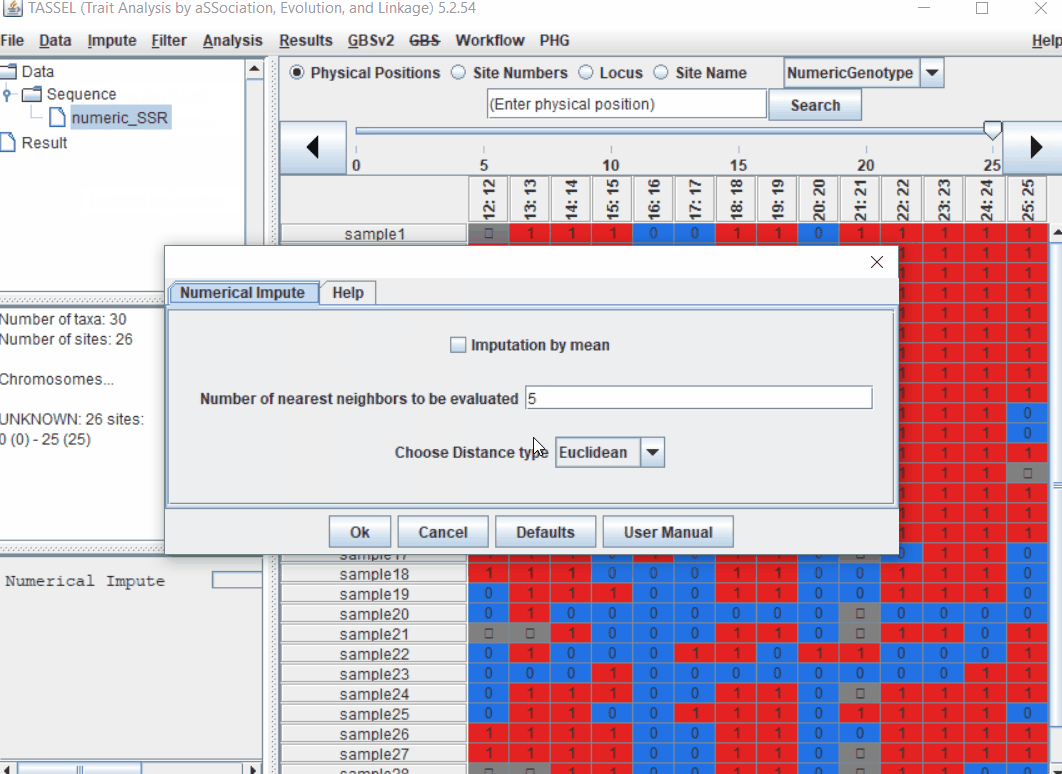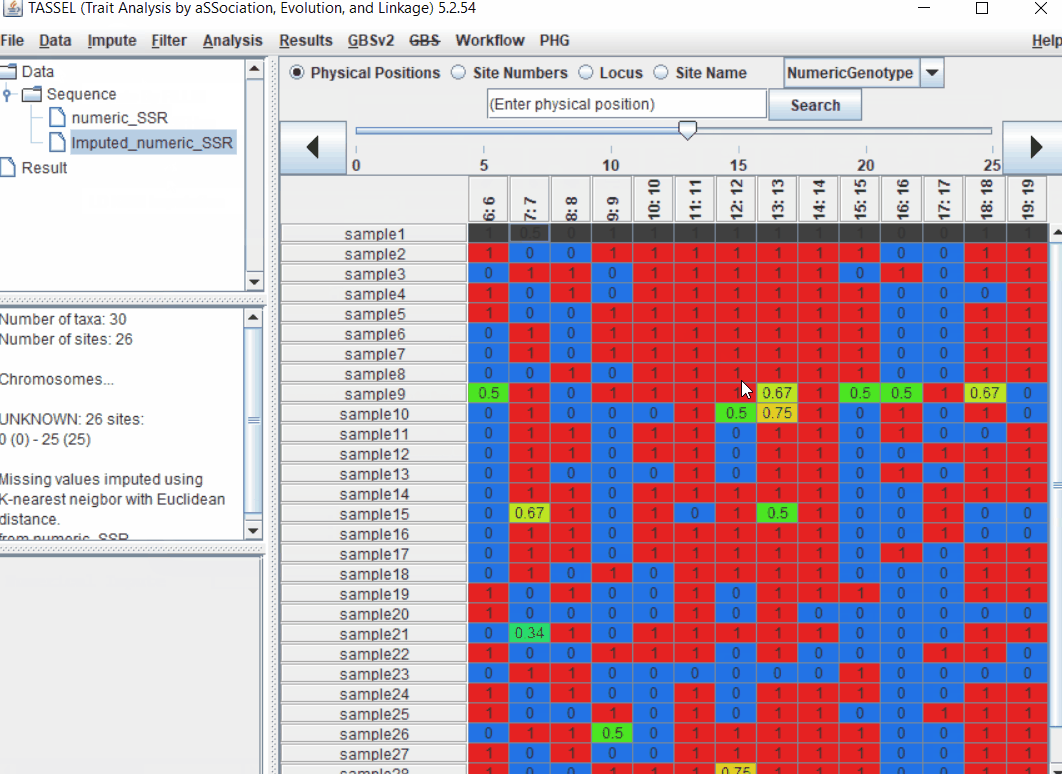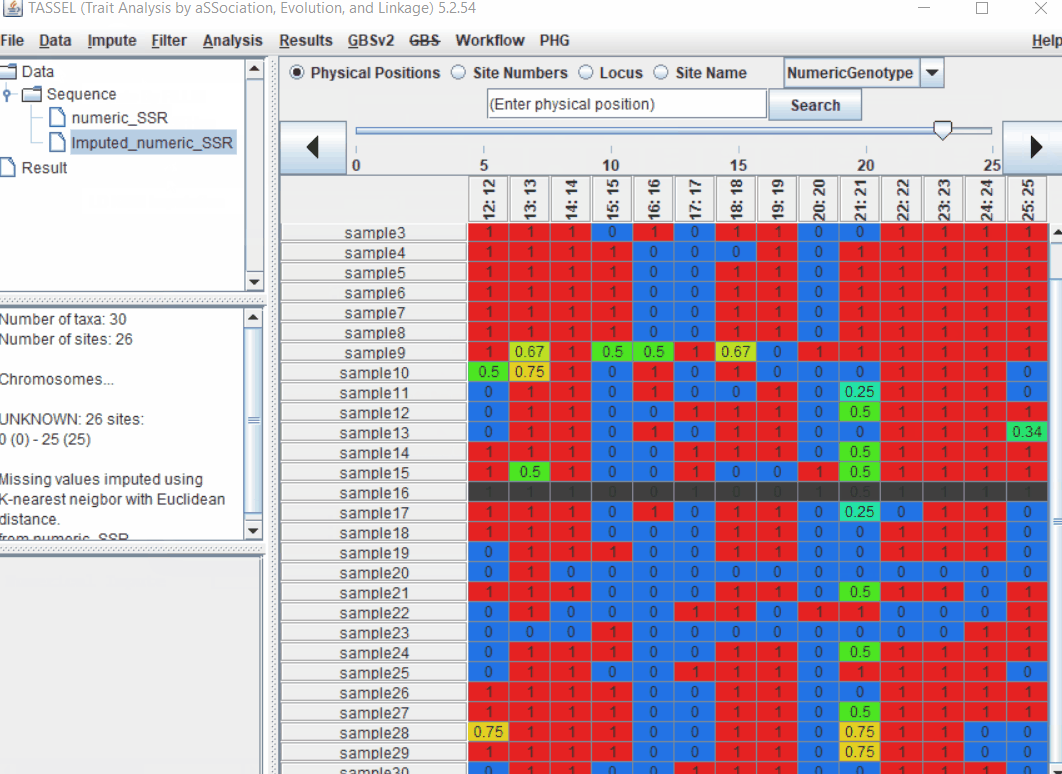
1.4 PCA analysis
Follow the below steps to perform PCA analysis:
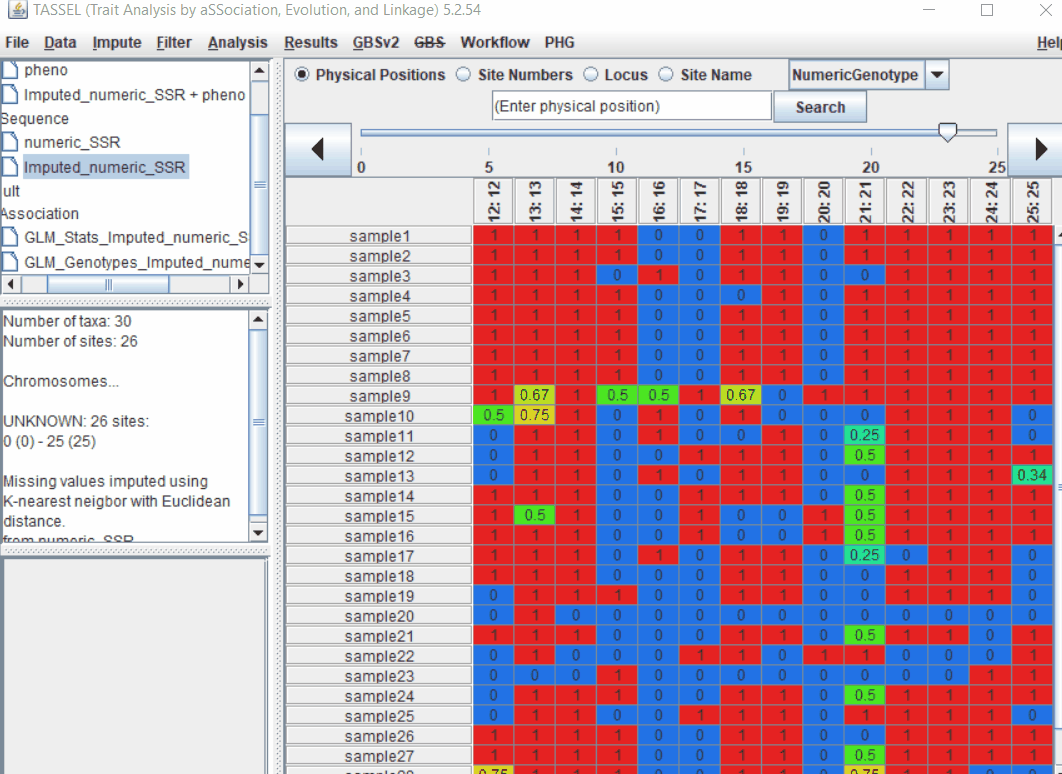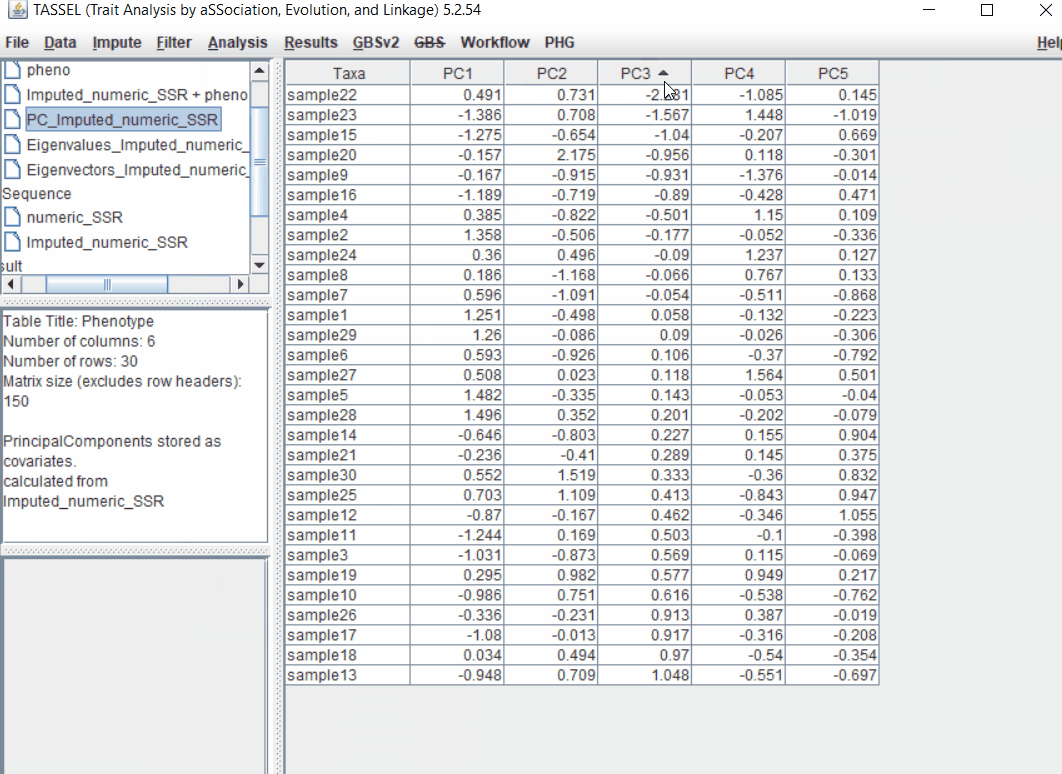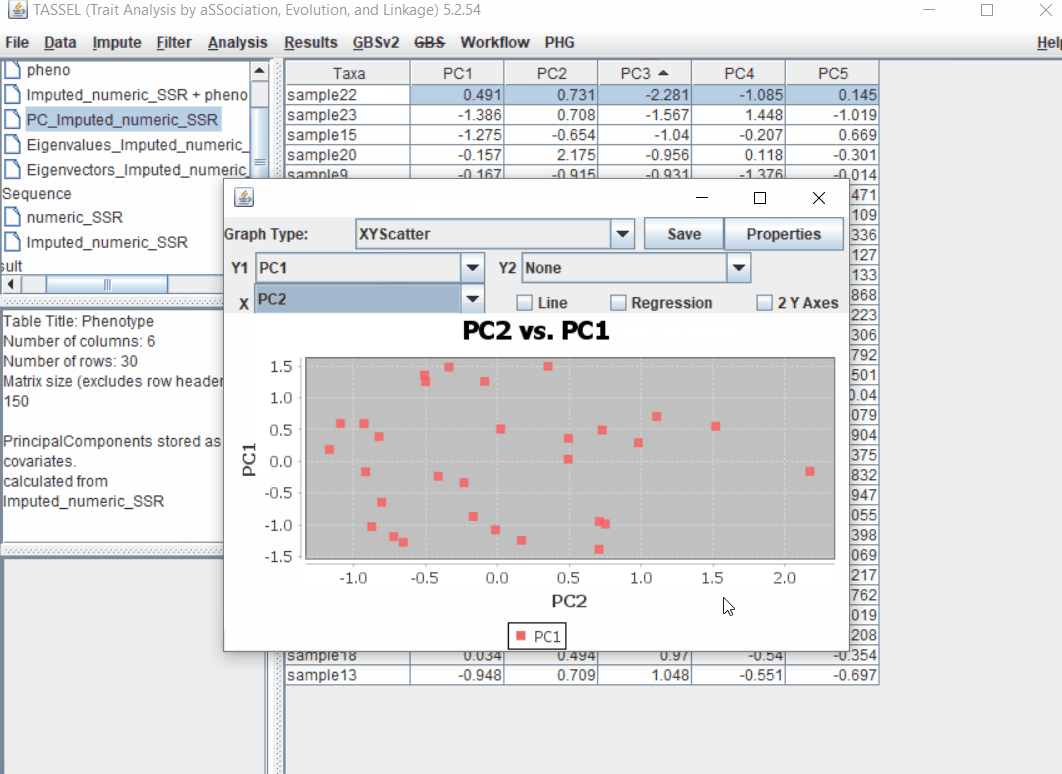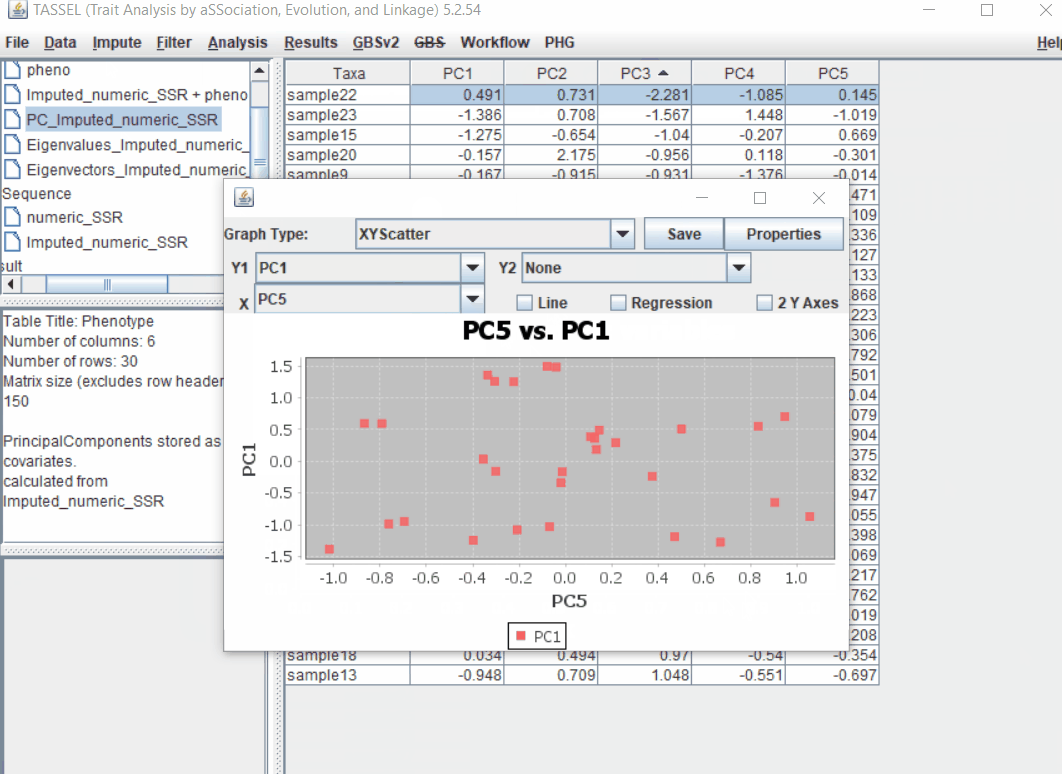
1.5 perform GLM
Run the GLM analysis by selecting the intersected files following the steps below:
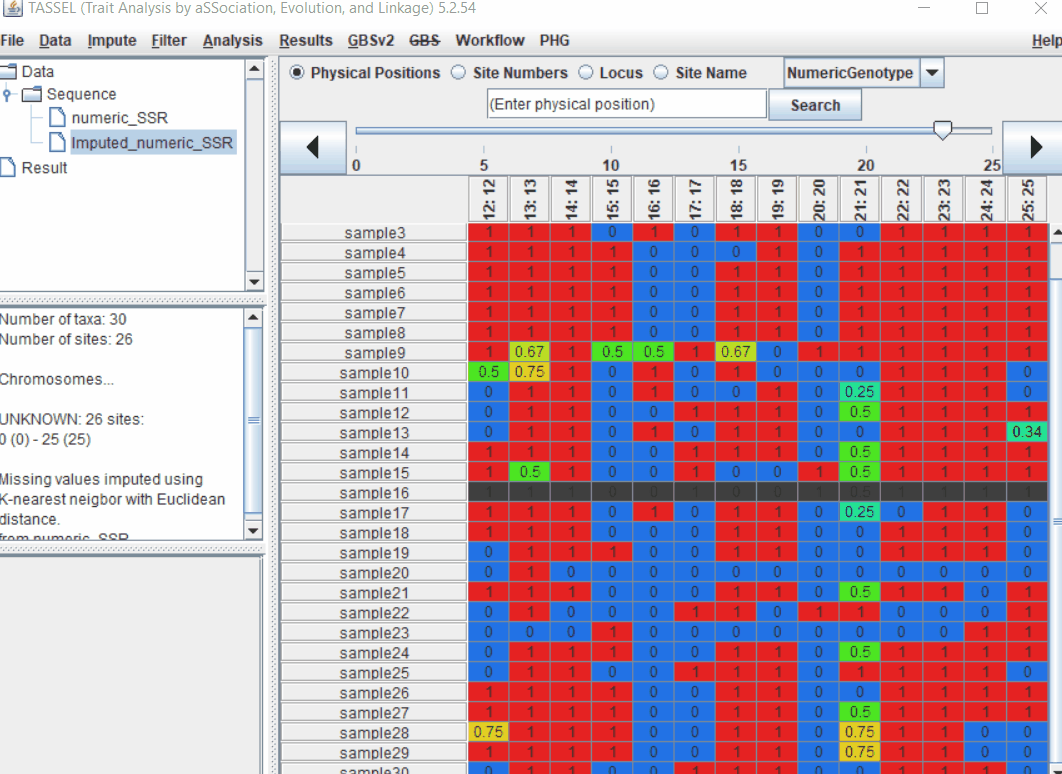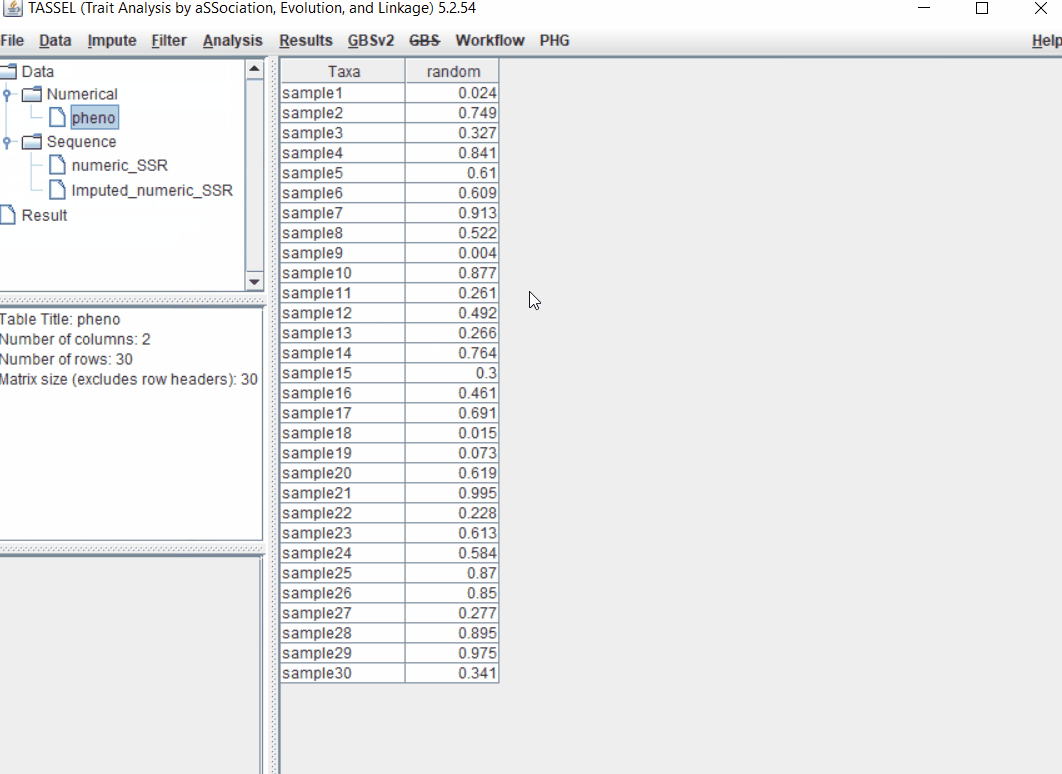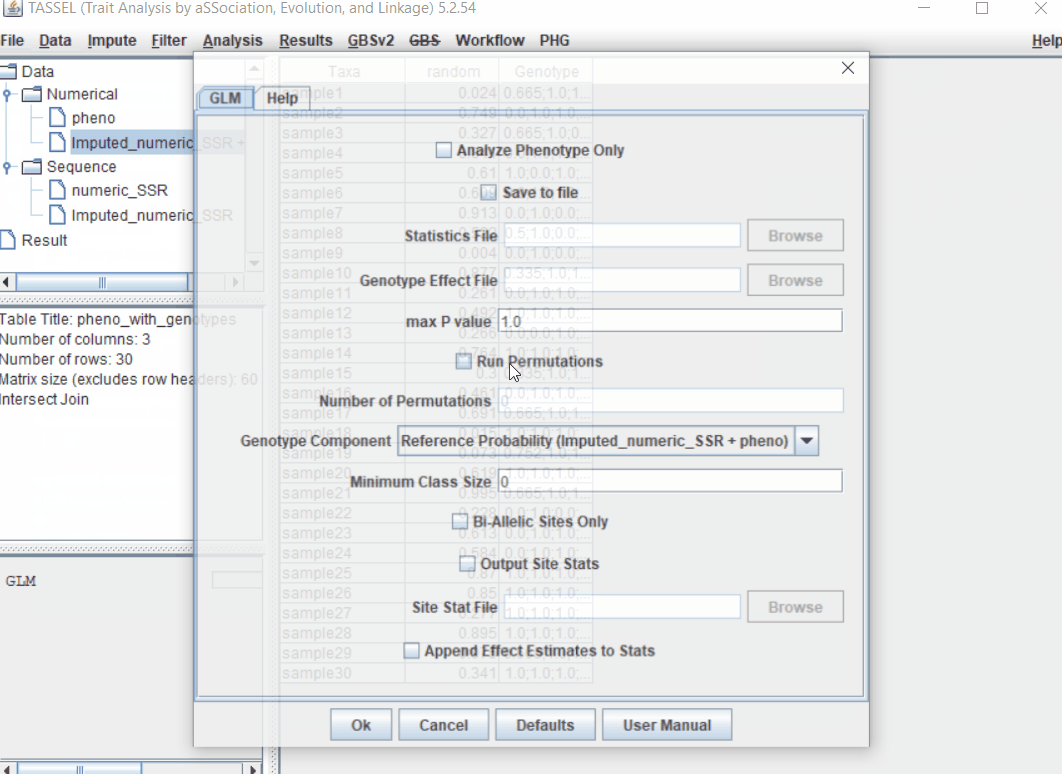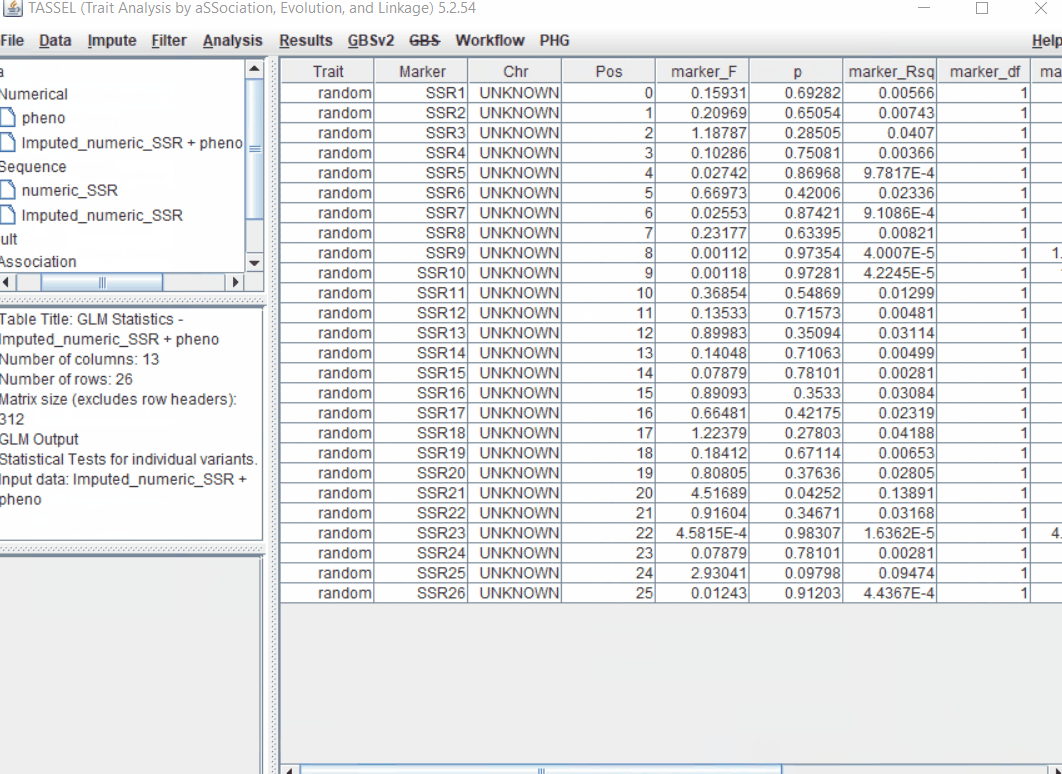
2.0 Convert Genotype calls into Numeric format in TASSEL
To convert your genotype data (A,C,G,T) into numeric format (0, 0.5, 1) follow the below steps:
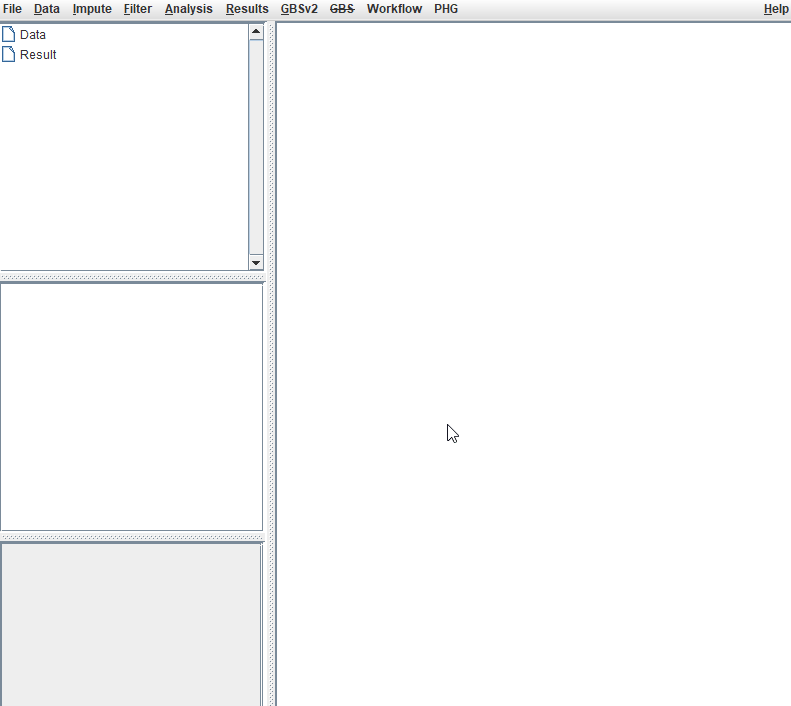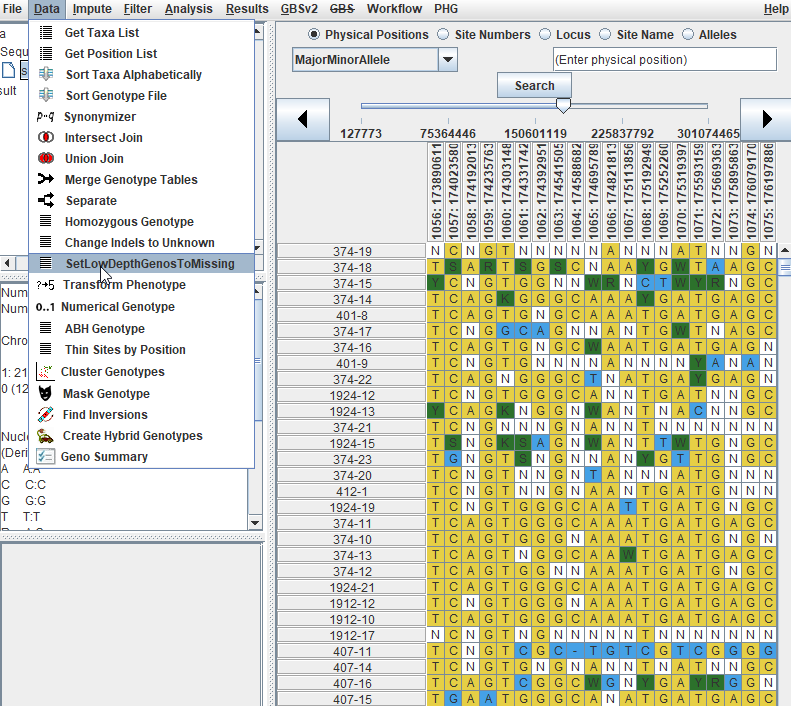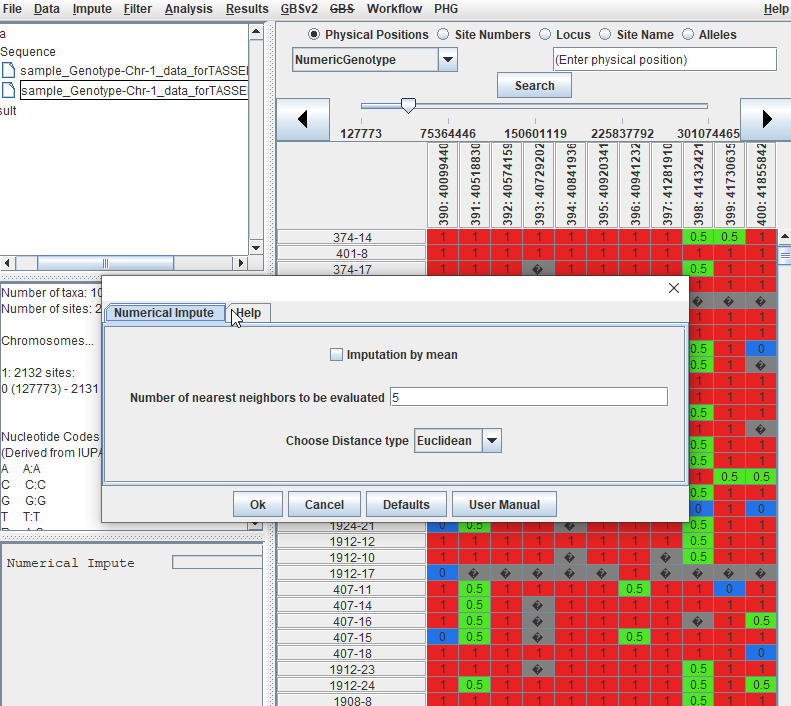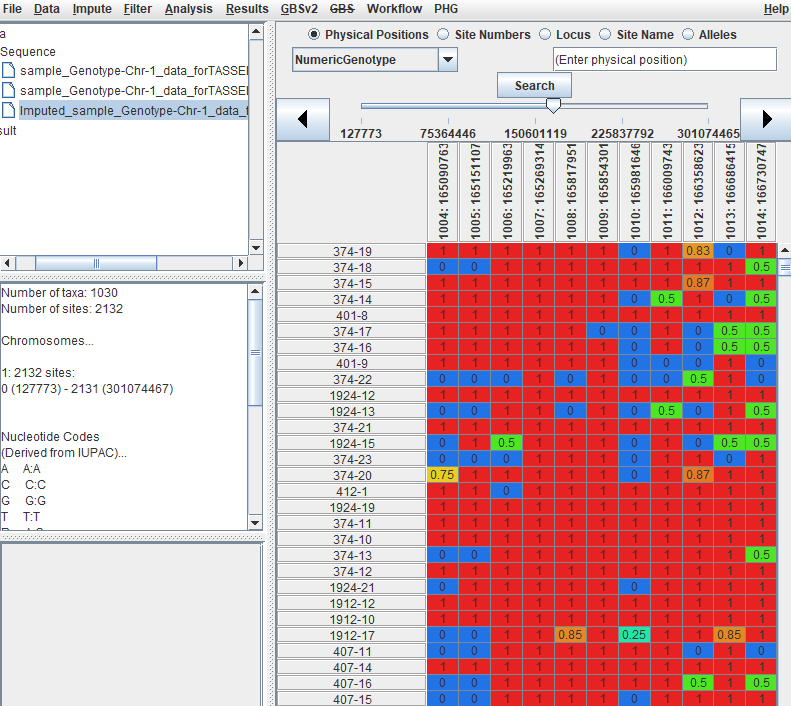
--- End of Tutorial ---
Thank you for reading this tutorial. If you have any questions or comments, please let me know in the comment section below or send me an email.
Bibliography
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES. (2007) TASSEL: Software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633-2635.